#How to trim a sequence in bioedit download#
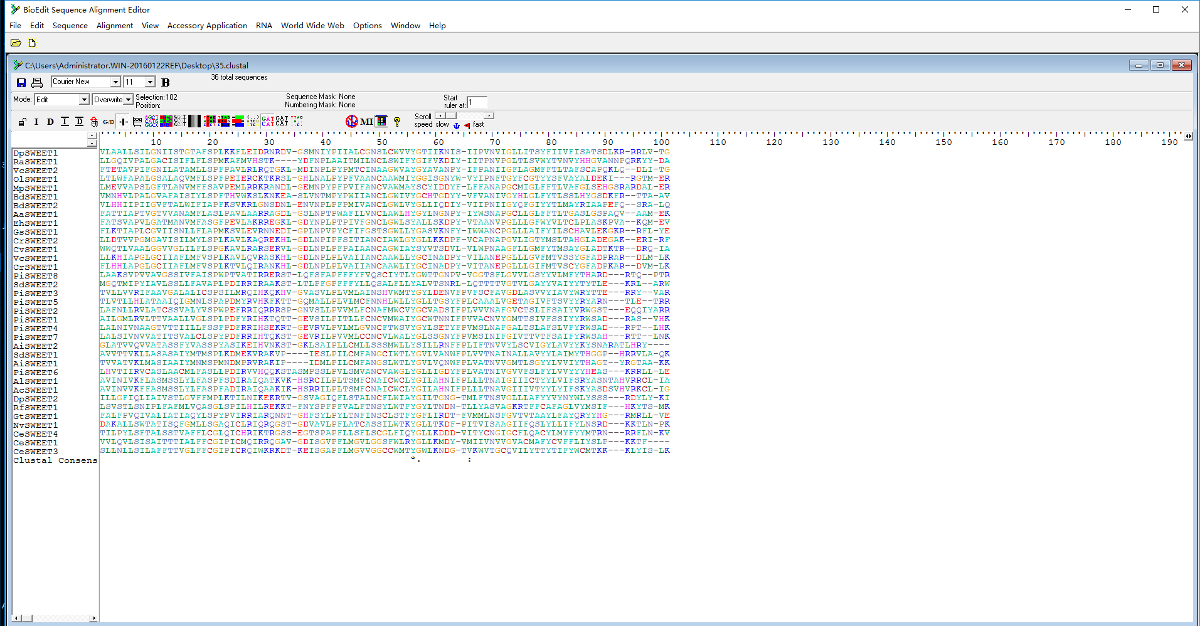
Or, you can download it first, then manipulate the data locally line command. Start with those under the group Text manipulation. There are many other tools to manipulate data within Galaxy if you need to rearrange before downloading. This will 'invert' your reverse sequence(s) so it (they) should now align with the forward sequence(s). You'll need to review the tool documentation/support to find out what data it works with. BioEdit is a sequence alignment editor and sequence analysis program that includes features such as Split window view, user defined color, information-based shading and auto integration with other programs such as ClustalW. I'm not sure about Bioedit - I haven't used it and the expected input formats are not clearly stated on the tool's website.
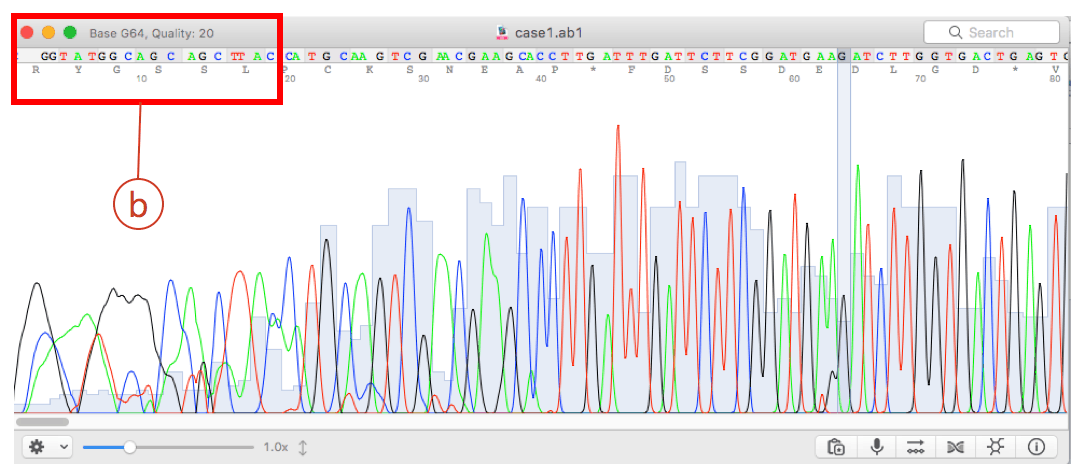
DNA chromatograms should be used to decide the final length. Interval is also a more specific version of tabular data.Įxcel will accept any tabular data as an input (with limits on the number of lines). You can use MEGA to trim sequences together at same length. Trim to Reference eliminates the ends of sequences that extend beyond an assembled Reference sequence. Or, you can convert SAM/BAM data to an interval datatype with the tool NGS: SAMtools: Convert SAM to interval. Trim Vector removes sequence-specific data contaminating the ends of your sequences. The SAM datatype is a version of tabular formatted data.
